1. Introduction
In the last century, scholars across a variety of fields have explored the mathematics of decision-making, or learning to intervene within a system. A challenge that relies on decision-making is fundamentally different, and more complex, than a classification problem. For example, designing a machine to distinguish a photo of a dog from a cat (a classification problem) is different from teaching one to recommend a photo of a dog versus a photo of a cat in order to maximize a key parameter (a decision-making problem).
Decision-making problems become even more complex when we consider that historical observed data result from past decisions, which may have been biased and/or do not convey the whole picture. For example, assume we have observed that carrying an umbrella is highly correlated with the likelihood that a car accident will occur. Can we recommend the intervention “do not carry an umbrella” in order to reduce the average number of accidents? Of course not, considering that carrying an umbrella and the car accidents are both outcomes prompted or caused by the rain.
In principle, a randomized test that is expected to disentangle the independence between the treatment and all baseline variables can solve this issue with biases in observed data. To take up our earlier example, if we carry an umbrella randomly regardless of the weather and then measure the accident rate, we will remove the effect of rain on the outcome.
However, there are many cases in which a randomized test is not feasible—such tests can be very expensive, and, in some circumstances, even unethical. Furthermore, even in an experimental setup, tests will not always go smoothly and thus may require statistical adjustments. One of the biggest challenges with randomized tests is the issue of compliance; for example, people who are assigned to perform (or not perform) a task might not follow the specified procedures. Causal Inference offers a variety of techniques that can help in cases in which either a randomized test is not feasible or the outcome of the test is not reliable.
At Wayfair, there are many business cases which face these challenges, and thus benefit from the use of causal inference techniques. In customer service, for example, we would like to measure the effect of various agents’ behaviors on customer satisfaction, which we assess through a survey sent after each contact. Politeness is one of those behaviors. Clearly, we can’t conduct any randomized tests to assess the impact of this behavior, because it is unethical and impractical to randomly be impolite to customers. Thus we only have access to historical observations to inform our assessments. In observation data, correlation is easy to measure, the challenge here is to estimate how much of the correlation is because of causality.
For this purpose, we need a third variable that explains both control and target variable. Reichenbach’s common cause principle, the basis of causal inference, asserts that this variable always exists. This principle states:
If two random variables X and Y are statistically dependent, then there exists a third variable Z that causally influences both variables. (In a special case Z might coincide with either X or Y) [1]
Though we may not always observe variable Z (which I will refer to as a set of covariates), Reichenbach’s principle claims that it exists, and this claim has been tested by experimental studies in many fields [2]. As I intend to show, the biggest challenge in causal inference is finding this set of covariates Z which influences both the treatment and outcome.
To illustrate the principle discussed above, in this blog post I will compare two known approaches that have been used across multiple domains in academia and industry:
1) Neyman-Rubin's Potential Outcome Theory (and its variations) 2) Pearlian do-calculus using the Bayesian Network framework
In this comparison, I will first evaluate the effectiveness of each approach and outline the particular challenges of using them. Then I will compare the two methods using synthetic data. The challenges encountered in employing each method should drive home the importance of finding the correct set of covariates to use when conducting causal inference.
2. Approach One: Neyman-Rubin's Potential Outcome Theory [3]
Assume we have to choose between two decisions (for example, yes/no or on/off). In Neyman-Rubin’s Potential Outcome Theory (referred to hereafter as “potential outcomes”), we design separate models for each of those decisions. To put it in mathematical terms, assume we have a set of variables x which has two components {x0 , x1}, referred to as control and treatment, respectively. Applying this to the umbrella example, we would say that x0 represents “not carrying an umbrella” and x1 represents “carrying an umbrella.” We would then take our observed data and split it into two buckets: x0 those who did not carry an umbrella, and x1 , those who carried an umbrella.
One of the common methods used in conjunction with potential outcomes is called the two model approach. With the two model approach, we try to separate the outcome variable Y into two models; we model Y0 for the control group and Y1 for the treatment group. The objective will be to approximate the “Average Treatment Effect (ATE)” Δ = E [Y1 - Y0]. Applying this to our umbrella example, Y represents whether or not an individual got into an accident. Here, we are looking to see which Z variable (with our umbrella example, daytime versus nighttime, city versus country environment, or perhaps, presence or absence of rain) accounts for the effect of umbrella carrying and getting into an accident.
In order to find the causal impact of X on the variable Y, we need to determine whether the control and target variables are a function of a third variable. Thanks to Reichenbach’s common cause principle, we know a set of variables Z exists that explains both X and Y; in order to model Y1 and Y0 we need a set of variables Z, such that at a given Z, the changes in input won’t change the target variables or X⊥Y0 , Y1 | Z (X and Y0 , Y1 are independent given Z). Applying this fact of independence the joint distribution will change as follows:
[latex]P(X, Y_0, Y_1| Z) = P (X|Z) P (Y_0, Y_1|Z)[/latex]
The biggest challenge with the potential outcomes approach lies in this assumption: X⊥Y0 , Y1 | Z. Our estimation of causal inference heavily depends on exactly how well this assumption holds, but unfortunately because we never observe Y0 and Y1 for the same sample, we cannot test the assumption. We are further limited in our evaluation by our insufficient knowledge of the system. The fact that this method does not provide any tools or instruction for detecting covariates Z correctly makes it vulnerable to error. The one condition that Rubin mentions in his paper (see [2]) is that no variable in Z should be chosen after the treatment is applied. In practice this is difficult, as it is sometimes hard to evaluate the relative timing of each variable when using observational data. I will show in section 5 that even if we select a set of variables Z ' that contains the correct covariates as a subset (i.e. Z ⊂ Z ') there is a chance that we will still not be able to predict the intervention correctly.
When implementing the two model approach as a part of potential outcomes, one can use multiple methods for constructing Y0 and Y1 using Z. The simplest way is to assume Y0 and Y1 are linear models of Z and that X ∈ {0,1} will not appear in Y0 . Other methods, including matching techniques, are also commonly used to calculate causal inference using potential outcomes. This can be achieved using various distance metrics including propensity matching, Mahalanobis matching, and genetic matching. I recommend taking a look at this amazing blog [4] written by Iain Barr for a deeper dive and better detailed explanation of various techniques. I have used some of his code in my analysis below (section 5) as it is written very clearly and it perfectly serves the purpose of this blog.
While there are ways to implement other methods to compensate for this approach’s lack of detection methods for the correct covariates, the fact that this common and popular method is particularly vulnerable to this failing emphasizes the extreme importance of finding the right covariates for causal inference.
3. Approach Two: Pearlian Do-Calculus
Now let’s examine another approach to causal inference. At the core of the Pearlian causal inference method is the construction of a network between our variables (this is the “Bayesian network”) and the definition of a set of variables called “back-door” variables. In the following section, I will focus on introducing these concepts and then we’ll see how they are connected to causal inference.
3.1 Bayesian networks as an I-Map (Map of Independence)
A Bayesian network is a directed acyclic graph, meaning the connections between nodes have directions and there are no cycles within the graph. It is a map that represents independence across a set of variables. In a Graph G, with {xi : i = 1, 2, ..., n} as the sets of variables, we can write the probability joint distribution as follows:
[latex]P(X_1, X_2, ..., X_n) = \prod_{i=1}^n P(X_i|pa_i)[/latex]
in which pai refers to the parent of xi.

Figure 1. An example of a Bayesian Network.
Let’s take a look at Fig. 1, in which the parents of node X1 includes pa1 = {X4 , X5}. The joint distribution based on the graph in Fig. 1 can be written as follows:
[latex]P(X_1, X_2, X_3, X_4, X_5) = P(X_4)P(X_5)P(X_1|X_4, X_5)P(X_3|X_1) P(X_2|X_1)[/latex]
In other words, each child-parent family in a graph G represents a deterministic function:
[latex]X_i = f_i(pa_i, \epsilon_i) (i = 1,...,n)[/latex]
in which ∈i (1 ≤ i ≤ n) are mutually independent, arbitrarily distributed random variables.
In a graph, a path that connects two variables is called a trail; a trail is active when X and Y are not independent given the connecting variables. If there is not a direct edge between X and Y, they are connected through a third variable Z by one of the following situations:

Figure 2: All possible combinations for three connected variables in a graph
A. Indirect Causal Effect (Fig. 2a): X → Z → Y The trail is active if and only if Z is not observed
B. Indirect evidential effect (Fig. 2b): X ← Z ← Y The trail is active if and only if Z is not observed
C. Common cause (Fig. 2c): X ← Z → Y The trail is active if and only if Z is not observed
D. Common effect (Fig. 2d): X → Z ← Y The trail is active if and only if either Z or one of Z’s descendants are observed.
As shown in Fig. 2, one can observe the independence and conditional independence between different variables by looking at a given Bayesian Network. In order to give greater form to concept, it is important to understand a few definitions along the way. First we must learn about “D-separation.” This concept will help us find more important variables which will make the target and the control variables independent.
D-Separation: Let X, Y, and Z be three sets of nodes in G. We say that X and Y are d-separated given Z if there is no active trail between any node x ∈ X and y ∈ Y given Z.
Next, we need to define “back-door variables.” This definition is so important, we will dedicate a whole section to it!
3.2 Back-door variables
We discussed the importance of the covariates in the previous sections. In Bayesian networks, we can detect the right covariates using the connections between variables. The following definition is one of the main types of variables that can be used for intervention calculation.
Back-door criterion: [5]: A set of variables Z satisfies the back-door criterion relative to an ordered pair of variables (Xi , Xj) in a directed acyclic graph G if: (i) no node in Z is a descendant of Xi, and (ii) Z blocks every active trail between Xi, and Xj via the parents of Xi

Figure 3. An example taken from [4] to explain the back-door criterion.
As an example, in Fig. 3, the following sets of variables are matching with back-door criteria for the sets of (Xi , Xj):
Z1 = {X1, X4} Z2 = {X1, X3, X4} Z3= {X4, X5} Z4 = {X1, X4, X5} Z5 = {X1, X3, X4, X5} Z6 = {X2, X4} Z7 = {X2, X4, X5} Z8 = {X3, X4} | Z9 = {X1, X2, X3, X4} Z11 = {X2, X3, X4} Z12 = {X2, X3, X4, X5} Z13 = {X1, X2, X4} Z14 = {X1, X2, X4, X5} Z15 = {X1, X2, X3, X4, X5} |
After identifying a set of back-door variables, we can easily calculate the intervention. The following theorem provides the mathematics of how everything comes together and shows the role of the back-door variables:
G-Formula Theorem: if a set of variables Z satisfies the back-door criterion relative to (X, Y) then: [latex]P(Y|do(X=x) = \sum_z P(Y|X=x, Z=z) P(Z=z)[/latex]
Basically, when we identify the back-door variables, we can calculate the intervention through the equation above (hereafter referred to as Eq. 1). I will show later that we only need one set of the back-door variables in order to calculate the intervention. In the case that we have multiple sets of back-door variables, the answer will be the same if we use any of the sets.
So, let’s apply the Pearlian do-calculus on our umbrella example. The Bayesian network would probably look like this:

Figure 4. The graph for the umbrella problem.
The variable Rain, is our back-door variable for the setting of umbrella as the control and accident as the target variable. Looking at Eq. 1 (of course we could answer with certainty if we had the actual data), Rain will be the z variable, Umbrella will be the x, and the probability of being in an accident will be y. Intuitively we know that umbrella and accident are independent variables so p (accident | umbrella, rain) will be completely explained by rain i.e. p (accident | umbrella, rain) = p (accident | rain) because umbrella ⊥ accident | rain.
As you can see, the Pearlian do-calculus model does provide methods for detecting the correct covariates, improving on the potential outcomes approach. However, it still presents challenges of its own which we will explore in our tests of these approaches below.
4. Making Synthetic Data
Now that we understand the pros and cons of these two common approaches, let’s see how they play out in practice. In order to put these theories to the test, first, I need to create some synthetic data. This approach will help me to control the distribution of both input and their relationship to the output of the system. I can also generate randomized tests for the control variable that I would like to test. Consequently, I can produce the data in which the inference is very different from the observation; most importantly, I will be able to know exactly how much the causal inference of my control variable is on the target.
I have used the following python packages to make this tutorial; most of them are standard python packages, “causalgraphicalmodels” and “causalinference” are two specific ones that I use for finding the back-door variables and also calculating the various potential outcomes model.
import numpy as np
import seaborn as sns
import itertools
import statsmodels.api as sm
from scipy.stats import norm
import causalgraphicalmodels as cgm
import causalinference as ci
Note: the following function must be added to causalgraphicalmodels/example.py
cie_example = StructuralCausalModel({
"z1": lambda n_samples: np.random.normal(size=n_samples),
"z2": lambda n_samples: np.random.normal(size=n_samples),
"z8": lambda n_samples: np.random.normal(size=n_samples),
"z9": lambda n_samples: np.random.binomial(1, p=0.01, size=n_samples) *\
np.random.normal(size=n_samples) +\
np.random.normal(loc=0.1, scale=0.01, size=n_samples),
"x": logistic_model(["z1", "z2"], [-1, 1]),
"z3": linear_model(["x"], [1]),
"y": linear_model(["z3", "z5", "z9"], [3,-1, 30]),
"z4": linear_model(["z2"], [-1]),
"z5": linear_model(["z4"], [2]),
"z6": linear_model(["y"], [0.7]),
"z7": linear_model(["y", "z2"], [1.3, 2.1])
})
After updating example.py, we have our data generator ready:
cie = cgm.examples.cie_example
cie_cgm = cie.cgm
cie_cgm.draw()

Figure 5. The Bayesian network that is used for synthetic data.
data = cie.sample(n_samples=100000)
data.head()

Table 1. A sample of the synthetic data.
Fig. 5 shows the Bayesian network that shows the connection of the variables. Table 1 just shows an example of the data. I sampled 100,000 times from the network shown in Fig. 4, equivalent to the number of rows of observation.
In order to simulate the randomized test, we can simply use the same network and remove all the arrows going into node x ; i.e. in our graph, the value of x is controlled by variable z2 but we would like to randomly change the value of x. Thus, by detaching the connection between z2 and x (removing the effect of z2), we can randomly assign values to our control variable x.
cie_cgm.do('x').draw()
Figure 6. The Bayesian Network for the randomized test.
As Fig. 6 illustrates above, the structure of a randomized test can be achieved by simply removing all of the connections from the variables (z1 and Z2) that control variable x.
randomized_test = (cie
.do('x')
.sample(n_samples=100000,
set_values={"x":
np.random.binomial(p=0.5, n=1, size=100000)}))
sns.distplot(randomized_test[randomized_test.x==1].y.clip(-25, 30), label='Treatment')
sns.distplot(data[data.x==1].y.clip(-25,30), label='Observation for Treatment')
sns.distplot(randomized_test[randomized_test.x==0].y.clip(-25, 30), label='Control')
sns.distplot(data[data.x==0].y.clip(-25,30), label='Observation for Control')
plt.ylabel('Probability Density')
plt.legend()

Figure 7. Probability density function for y conditioned on x ∈ {Treatment=1, Control=0} for both randomized tests and observations.
As shown in Fig. 7, there is a slight difference in the uplift of variable in x within observational and the results of the randomized test. The actual difference can be calculated as follows:
ATE = randomized_test[randomized_test.x==1].y.mean() - randomized_test[randomized_test.x==0].y.mean()
ATE_obs = data[data.x==1].y.mean() - data[data.x==0].y.mean()
print(f"Actual ATE:{ATE}")
print(f"Observed ATE: {ATE_obs}")

The observation estimates the value of the intervention almost 50% wrong (48%). In the following sections, I will use various techniques to estimate the “Actual ATE” using only observational data.
5. Testing the Potential Outcomes Method
So let’s put the potential outcomes theory to the test. We just generated the data and we are ready to see how far we can go with this approach. As outlined in the description of this theory above, potential outcomes can be a very powerful approach if one knows the correct covariates. I will estimate the ATE within the potential outcome framework mostly using two model approaches using various sets of covariates. The first set I will try with all available observed covariates in my data. This practice will show that even if the correct covariates are a subset of the selected variables, the estimation still will be unsuccessful!
For the second set, I will choose the right set of covariates. I will do this by borrowing from Pearlian do-calculus and choosing from the back-door variable sets. In this exercise, I am trying to show that when the covariates are correctly selected, all the variations of modeling in potential outcomes converge to the same solution. Although this cannot be generalized to every problem, I would like to emphasize the importance of choosing the right covariates compared to making a more complex model.
5.1. All variables as the covariates
zs = [c for c in data.columns if c.startswith("z")]
cm = ci.CausalModel(Y=data.y.values,
D=data.x.values,
X=data[zs].values)
cm.est_via_ols()
cm.est_via_matching()
cm.est_propensity_s()
cm.est_via_weighting()cm.stratify_s()
cm.est_via_blocking()print(cm.estimates)

y = []
yerr = []
x_label = []for method, result in dict(cm.estimates).items():
y.append(result["ate"])
yerr.append(result["ate_se"])
x_label.append(method)x = np.arange(len(y))
plt.errorbar(x=x, y=y, yerr=yerr, linestyle="none", capsize=5, marker="o")
plt.xticks(x, x_label)
plt.title("Estimated Effect Size", fontsize=18)
plt.hlines(ATE, -0.5, 3.5, linestyles="dashed")
plt.xlim(-0.5,3.5);

Figure 8. The dashed line shows the actual ATE from a randomized test.
The estimation of the ATE using various techniques and all the variables at the covariates is shown in Fig. 8; in this figure the dash line is the actual value. The results clearly show how off the mark the estimation is, and how these results didn’t substantially improve when we introduced more complex modelling techniques to our calculations.
5.2. Z2 as the covariates
I will show in the next section how we can choose the right covariates using the back-door variable definition, but for now assume that I magically select Z2 as the covariates.
NOTE: the code is pretty much the same for generating the outputs the only change is that the variable “zs” must be replaced as follows:
zs = ['z2']


Figure 9. The dashed line shows the actual ATE from a randomized test.
The results shown in Fig. 9 confirm our hypothesis regarding the importance of the covariates versus the complexity of the models. The graph shows that the true value of the actual ATE is within the range of the estimations of the four methods with which we experiment. In this case, the complexity (nonlinearity) of the methods did not add any value to the accuracy of the final output.
This test suggests that the potential outcomes approach is an effective technique which provides different methods to estimate the causal inference. The only challenge is that potential outcomes does not provide a clear instruction for choosing the right the covariates.
6. Testing the Pearlian Do-Calculus Approach
6.1. List of back-door variables
In the potential outcomes section, we discovered that choosing the covariates correctly is a crucial step. Bayesian networks provide the tools and facilitate this process through the selection of so-called ‘back-door variables.’
Referring to section 3.2, the “causalgraphicalmodels” package in python provides the list of back-door variables for a set of control-target variables for a given graph.
cie_cgm.get_all_backdoor_adjustment_sets("x", "y")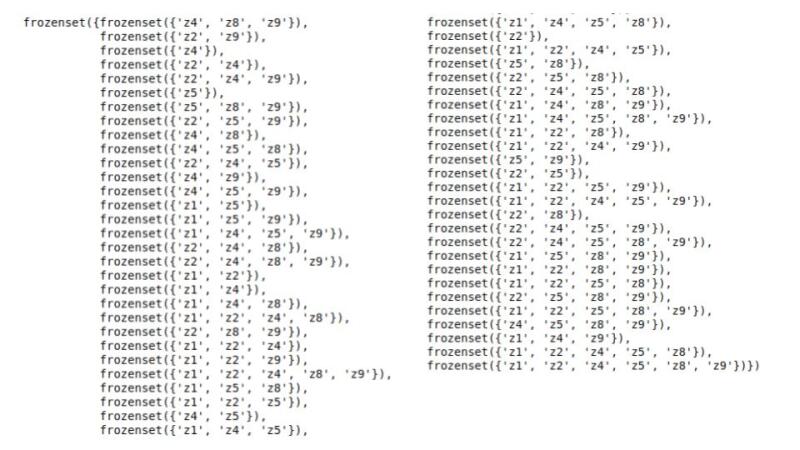
I have presented all the back-door variable sets above; you can use these as an exercise to to test if you can find the same list or at least can find the rationale behind each of these sets.
If we choose any set from this list, potential outcomes can estimate the ATE correctly (I have shown this with only one set).
6.2. Using Eq. 1 for calculating the ATE
Eq. 1 (copied below for convenience) requires the joint distribution of a set of covariates Z. We also need the P (Y|X = x, Z = z) which is basically a function that maps Z and x to Y.
In our case, Z = {Z2} which simplifies the joint distribution to a single probability density function; I will make two OLS models with Z2 separately for X = 0 and X = 1.
[latex]P(Y|do(X=x) = \sum_z P(Y|X=x, Z=z) P(Z=z)[/latex]
Let's make the first OLS for the observational data conditioned on X = 1.
# The X_ is the covariates while X==1
Z_ = data[data[X]==1]['z2']
Y_ = data[data[X]==1]['y']# Add a constant in order to have an intercept in OLS
Z_ = sm.add_constant(Z_)# Fitting the OLS model
y_x1_z = sm.OLS(Y_, Z_)
y_x1_z_model = y_x1_z.fit()
I will make the same OLS for X = 0
Z_ = data[data.x==0]['z2']
Y_ = data[data.x==0]['y']
Z_ = sm.add_constant(Z_)
y_x0_z = sm.OLS(Y_, Z_)
y_x0_z_model = y_x0_z.fit()
In order to estimate the P (Z2), I assume a normal distribution with the µ , σ calculated from the observation. Of course, in more general cases one should calculate P (Z) using more elaborate techniques.
mu = data[Z].mean()
std = data[Z].std()
z_ = data[Z].values
p_z = lambda z: norm.pdf(z, mu, std)
So we have all the ingredients ready! Let’s calculate the P (Y |do (X = x))
# probably the better practice is to estimate the Y standard deviation as a
# function of z2. For now I assume the std won't change from observation to
# intervention (which is not the case)
y_std = data[Y].std()
def p_y_x_z(z, y, model):
return norm.pdf(y, model.predict([1, z]),
int(y_std)) * p_z(z)
def p_y_dox(ys, x):
if x==1:
model = y_x1_z_model
elif x==0:
model = y_x0_z_model
return np.sum(v_p_z(z_, ys, model))# np.vectorize helps me to pass a numpy array to a function that receives scalar
v_p_z= np.vectorize(p_y_x_z)
v_p_y_dox = np.vectorize(p_y_dox)# Because I'd like to calculate the whole density function of P(Y|do(x)) I have to
# calculate the Eq(1) for different y. For calculating the ATE I don't need
# to save all of these values.yy = np.linspace(-20, 30, 60)
py_dox1 = v_p_y_dox(yy, x=1)
py_dox0 = v_p_y_dox(yy, x=0)
By using Z2 as the covariates:


Figure 10. The estimation of probability distribution of treatment and control effect using Z2 as the covariates.
By using Z5 as the covariates:


Figure 11. The estimation of probability distribution of treatment and control effect using Z5 as the covariates.
There are two important points illustrated in Fig. 10 and Fig. 11. First, choosing the right covariates can estimate not only the ATT or ATC, but also it can predict the whole probability distribution of the inference. Second, we have used two completely different sets of variables and as far as they fit the back-door variable criteria the results will be the same.
6.3. Learning the structure of Bayesian Networks
As we have seen so far, as opposed to the potential outcomes method, the do-calculus approach provides a powerful tool to find the relative covariates for estimating the intervention. The question remains as to how difficult it will be to estimate the network architecture, which is the crucial step for finding the “back-door” or “front-door” variables. Luckily, many structural learning algorithms have been created for this purpose. Providing a deep review of these important tools is unfortunately out of the scope of this blog. As such, I would like to show an example of an existing package in R called bnlearn that has implemented most of those algorithms efficiently, is open source, and it is relatively easy to use.
The equivalent in Python package is called
pgmpy, however the network learning modules in this package are much slower than those in bnlearn. So, for now I will try to show that technically it is possible to get the network from the observed data using bnlearn.
# ryp2 helps to run R modules in python
from rpy2.robjects.packages import importr
import rpy2.robjects.packages as rpackages
import rpy2
# importing bnlean
bnlearn = rpackages.importr('bnlearn')
readr = rpackages.importr('readr')
# importing the data that we generated in the previous section as csv
dtrain = readr.read_csv('data.csv')# Hill Climbing is the most common algorithm for structural learning
dag = bnlearn.hc(dtrain)
print(dag)
Bayesian network learned via Score-based methods:
Model:
[z9][z8][z2][z1][z4|z2][x|z2:z1][z5|z4][z3|x][y|z9:z5:z3][z6|y][z7|z2:y]
nodes: 12
arcs: 11
undirected arcs: 0
directed arcs: 11
average markov blanket size: 2.67
average neighbourhood size: 1.83
average branching factor: 0.92
learning algorithm: Hill-Climbing
score: BIC (Gauss.)
penalization coefficient: 5.756463
tests used in the learning procedure: 187
optimized: TRUE
In this example, bnlearn manages to learn the structure perfectly even though our data includes discrete and continuous variables which technically is called a hybrid network.
Conclusion
In this blog I have tried to show the challenges inherent in causal inference calculation, which ultimately relies on finding the right covariates. The potential outcomes approach provides better tools for calculating causal inference, but it does not provide a suitable path to find the right sets of covariates. On the other hand, the Pearlian do-calculus method can provide (at least in our example) the right tool for estimating the correct sets of variables, but presents a challenge in determining the graph structure.
One assumption I have made is that we have measured or observed the confounding variables within our data, which is not always the case. I will dedicate my next blog to the estimation of causal inference in situations where we have not observed the confounding variable and must approximate it first.
References
- Reichenbach, Hans. The direction of time. Vol. 65. Univ of California Press, 1956.
- Peters, Jonas, Dominik Janzing, and Bernhard Schölkopf. Elements of causal inference: foundations and learning algorithms. MIT press, 2017.
- Rubin, D. B. (2005). Causal inference using potential outcomes: Design, modeling, decisions. Journal of the American Statistical Association, 100(469), 322-331.
- Barr, Ian. Causal Inference With Python, http://www.degeneratestate.org/posts/2018/Mar/24/causal-inference-with-python-part-1-potential-outcomes/
- Pearl, J. (1995). Causal diagrams for empirical research. Biometrika, 82(4), 669-688.

